Adaptive search
RNAsubopt (part of the ViennaRNA package), in its
Boltzman sampling nature, usually produces only structures very near to the global minimum
over and over again. Generating samples with higher energy can be done by introducing a
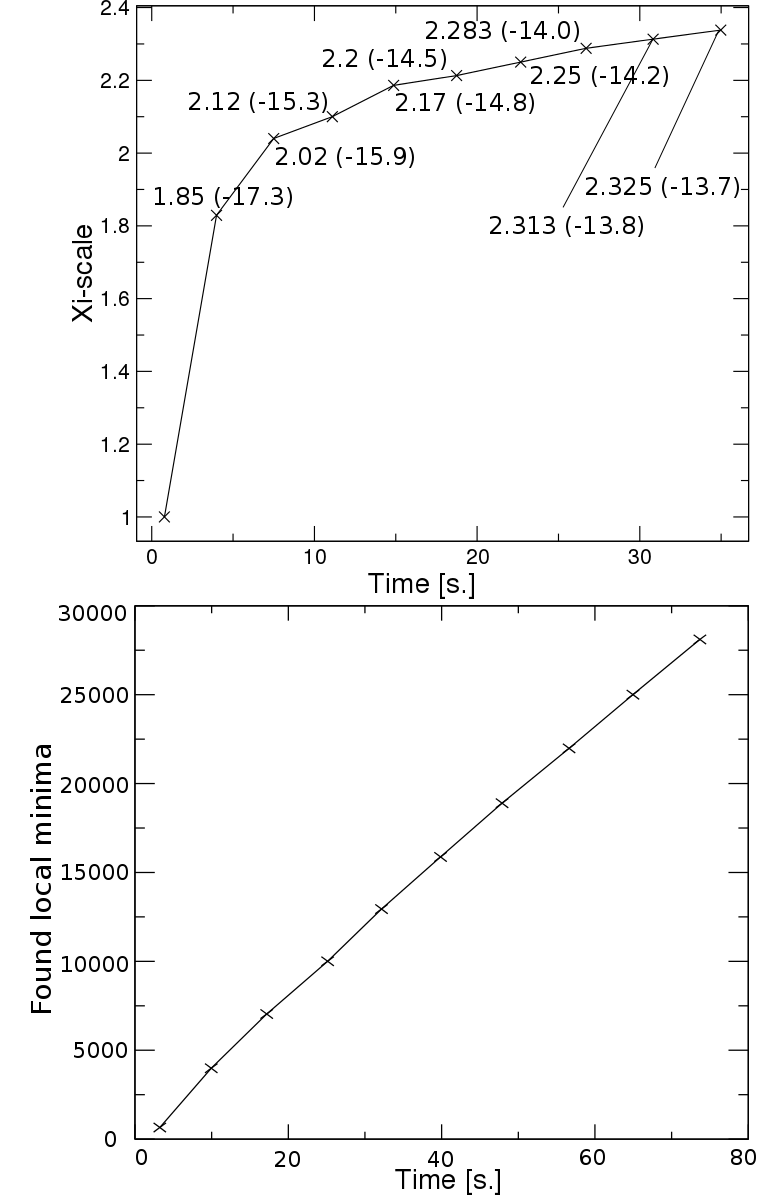
scaling parameter (xi) in the sampling, which can be seen as an alteration of the sampling
temperature. The adaptive search schedule aims to find local minima, that cover a desired
energy range from the global optimum. For this purpose, a criterion of coverage has been
introduced to be sure that we have successfully covered local minima up to some energy.
After this, sampling shifts to higher energies by increasing the scaling parameter, as
long as the desired energy range is not covered or a maximal number of iterations is not
reached.
The adaptive search schedule script is written in Python 2.7 and requires both programs
RNAsubopt and RNAlocmin to work. To make it run, just download
the script asearch_2.py, edit the paths to programs on the top
of the script, and run
python asearch_2.py -I input_fileThe output is generated in the same directory as the input file with filename extensions
.lm (basic output - list of generated local minima), .lmns
(list of local minima after filtering), and .bhg (BHGgraph if --bhg
option was specified). Run
python asearch_2.py -hto get help and available options.
First, it constructs xi-to-energy table, where the average energy values for different
scale parameters are stored. (New version constructs this table already on the run.)
Then it iteratively runs RNAsubopt and RNAlocmin and vary
the scaling parameter to obtain the best set of local minima.
The coverage criterion can be modified with a coverage parameter, which is by default
set to 0.1. Redefining it to a higher number shifts the sampling to higher energies much
sooner, and vice versa. We recommend using values between 0.05 to 0.8. You can find more
information in the article below.
In case you are using our software for your publications you may want to cite:
Basin Hopping Graph: A computational framework to characterize RNA folding landscapes
Bioinformatics 2014, doi:10.1093/bioinformatics/btu156, Supplementary material, ( bibtex)
Pseudoknots in RNA folding landscapes
Bioinformatics 2015, doi:10.1093/bioinformatics/btv572, Supplementary material, ( bibtex)
News
-
11 - 05 - 2015
Updated the asearch_2.py script. Changed the usage explanation to better capture the new version of the script.
-
22 - 10- 2014
New version of the adaptive searching script. Added support for pseudoknots and help.
-
24 - 03 - 2014
Added some clarification for usage of the asearch.py script.
Download
 Get the Python scripts
Get the Python scripts
Just download the python script below.
asearch_2.py (py, Python script) [19.2kB]
Older versions:
There should rarely be a good reason to use any but the latest version of our software. However if you want to look up the old bugs, here's a list with most of the older releases for download.
Contact
In case of problems, found bugs, or need to chat, do not hesitate to contact us:
- marcel
 tbi.univie.ac.at -- programming
tbi.univie.ac.at -- programming - qin
 bioinf.uni-leipzig.de -- ideas
bioinf.uni-leipzig.de -- ideas
Comments and bug reports should be sent to rna@tbi.univie.ac.at



