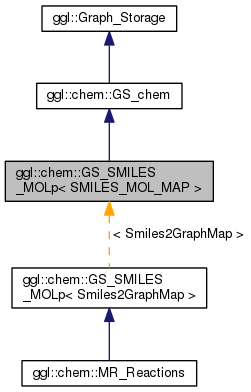
SMILES graph storage using STL SMILES-GraphPointer map. More...
#include <GS_SMILES.hh>
Public Member Functions | |
| virtual void | add (const Molecule &graph) |
| virtual void | addMolecule (const Molecule &graph) |
| GS_SMILES_MOLp (SMILES_MOL_MAP &smiles2mol) | |
| GS_SMILES_MOLp (SMILES_MOL_MAP &smiles2mol, const SMILES_MOL_MAP &smiles2molCheckOnly) | |
| GS_SMILES_MOLp (SMILES_MOL_MAP &smiles2mol, const SMILES_MOL_MAP &smiles2molCheckOnly1, const SMILES_MOL_MAP &smiles2molCheckOnly2) | |
| virtual | ~GS_SMILES_MOLp () |
Protected Types | |
| typedef boost::filtered_graph < Molecule, edge_is_in_component, node_is_in_component > | ComponentGraph |
| typedef std::vector< int > | ComponentIdVec |
| typedef boost::property_map < Molecule, PropNodeIndex > ::const_type | IndexMap |
| the type of a constant index map of a Molecule boost graph More... | |
Protected Member Functions | |
| virtual bool | insert2map (const std::string &SMILES, const Molecule &graph) |
Protected Attributes | |
| SMILES_MOL_MAP * | smiles2mol |
| const SMILES_MOL_MAP * | smiles2molCheckOnly1 |
| const SMILES_MOL_MAP * | smiles2molCheckOnly2 |
A Graph_Storage implementation that converts each added Molecule graph into a SMILES string representation and adds it, if not already existing, to the specified STL map container using the SMILES as key and the pointer to the newly allocated Molecule object as value.
| STL_INSERTER | an STL insert iterator (e.g. std::insert_iterator) to add all SMILES to store to |
Definition at line 180 of file GS_SMILES.hh.
|
protectedinherited |
a component graph definition if more than one connected component present is present and it is needed to split the graph to report into its components
Definition at line 128 of file GS_chem.hh.
|
protectedinherited |
the container type used to represent the connected component id for each node
Definition at line 32 of file GS_chem.hh.
|
protectedinherited |
Definition at line 35 of file GS_chem.hh.
| ggl::chem::GS_SMILES_MOLp< SMILES_MOL_MAP >::GS_SMILES_MOLp | ( | SMILES_MOL_MAP & | smiles2mol | ) |
Construction
| smiles2mol | the map (SMILES->molecule) to that each generated SMILES and its molecule is added to if not already present |
| ggl::chem::GS_SMILES_MOLp< SMILES_MOL_MAP >::GS_SMILES_MOLp | ( | SMILES_MOL_MAP & | smiles2mol, |
| const SMILES_MOL_MAP & | smiles2molCheckOnly | ||
| ) |
Construction
| smiles2mol | the map (SMILES->molecule) to that each generated SMILES and its molecule is added to if not already present |
| smiles2molCheckOnly | an additional map to check for existence of a new molecule to store |
| ggl::chem::GS_SMILES_MOLp< SMILES_MOL_MAP >::GS_SMILES_MOLp | ( | SMILES_MOL_MAP & | smiles2mol, |
| const SMILES_MOL_MAP & | smiles2molCheckOnly1, | ||
| const SMILES_MOL_MAP & | smiles2molCheckOnly2 | ||
| ) |
Construction
| smiles2mol | the map (SMILES->molecule) to that each generated SMILES and its molecule is added to if not already present |
| smiles2molCheckOnly1 | an additional map to check for existence of a new molecule to store |
| smiles2molCheckOnly2 | an additional map to check for existence of a new molecule to store |
|
virtual |
|
virtualinherited |
The reported graphs is split into its individual independent components. Each component is forwarded to addMolecule to be implemented by any sub class.
| graph | the graph object to add that encodes one or more molecules |
Implements ggl::Graph_Storage.
|
virtual |
Converts a given molecule graph to SMILES and adds it to the storage container.
| graph | the Graph object to add. |
Implements ggl::chem::GS_chem.
|
protectedvirtual |
Reimplemented in ggl::chem::MR_Reactions.
|
protected |
the map where each SMILES string is mapped to the represented molecule
Definition at line 186 of file GS_SMILES.hh.
|
protected |
Definition at line 187 of file GS_SMILES.hh.
|
protected |
Definition at line 188 of file GS_SMILES.hh.