Reaction match reporter. More...
#include <MR_Reactions.hh>
Public Types | |
| typedef std::set< Reaction > | Reaction_Container |
| a container that stores Reaction objects More... | |
Public Member Functions | |
| virtual void | add (const Molecule &graph) |
| virtual void | addMolecule (const Molecule &graph) |
| MR_Reactions (const Smiles2GraphMap &smiles2graph, Smiles2GraphMap &newSmiles2graph, Reaction_Container &reactions, const bool addEachComponent=false, const ReactionRateCalculation *rateCalculation=NULL, const AromaticityPerception &aromaticity=AP_NSPDK("Marvin:general:2013")) | |
| MR_Reactions (const Smiles2GraphMap &smiles2graph, const Smiles2GraphMap &smiles2graph2, Smiles2GraphMap &newSmiles2graph, Reaction_Container &reactions, const bool addEachComponent=false, const ReactionRateCalculation *rateCalculation=NULL, const AromaticityPerception &aromaticity=AP_NSPDK("Marvin:general:2013")) | |
| virtual void | reportHit (const sgm::Pattern_Interface &pattern, const sgm::Graph_Interface &target, const sgm::Match &match) |
| virtual | ~MR_Reactions () |
Protected Types | |
| typedef boost::filtered_graph < Molecule, edge_is_in_component, node_is_in_component > | ComponentGraph |
| typedef std::vector< int > | ComponentIdVec |
| typedef boost::property_map < Molecule, PropNodeIndex > ::const_type | IndexMap |
| the type of a constant index map of a Molecule boost graph More... | |
| typedef SMILESwriter | Smiler |
| the SMILESwriter to use for target-to-smiles conversion More... | |
| typedef GS_SMILES_MOLp < Smiles2GraphMap > | SuperClass |
| access to the superclass of this class More... | |
Protected Member Functions | |
| virtual bool | insert2map (const std::string &SMILES, const Molecule &molGraph) |
Protected Attributes | |
| const bool | addEachComponent |
| Reaction | curReaction |
| the current Reaction to fill and to store later More... | |
| Graph2SmilesMap | graph2smiles |
| A vice versa lookup table for smiles2graph. More... | |
| GS_MolCheck | molchecker |
| const ReactionRateCalculation * | rateCalculation |
| if != NULL, for each new reaction a rate is calculated More... | |
| Reaction_Container & | reactions |
| The container of Reaction objects to fill and extend. More... | |
| ggl::MR_ApplyRule | Ruler |
| Smiles2GraphMap * | smiles2mol |
| const Smiles2GraphMap * | smiles2molCheckOnly1 |
| const Smiles2GraphMap * | smiles2molCheckOnly2 |
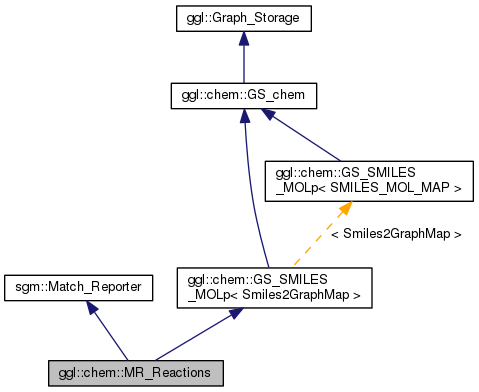
A Match_Reporter that generates a Reaction object from a rule
application.
NOTE : this class currently only works for target graphs that are an
object of Molecule_GraphV !!! Furthermore, it assumes that the
elements (pointer) from the covered target graph are from the here
maintained storage, to enable a fast lookup of their SMILES !!!
Finaly, SMILES are generated using an instance of SMILESwriter !!!
Definition at line 96 of file MR_Reactions.hh.
|
protectedinherited |
a component graph definition if more than one connected component present is present and it is needed to split the graph to report into its components
Definition at line 128 of file GS_chem.hh.
|
protectedinherited |
the container type used to represent the connected component id for each node
Definition at line 32 of file GS_chem.hh.
|
protectedinherited |
Definition at line 35 of file GS_chem.hh.
| typedef std::set< Reaction > ggl::chem::MR_Reactions::Reaction_Container |
Definition at line 104 of file MR_Reactions.hh.
|
protected |
Definition at line 109 of file MR_Reactions.hh.
|
protected |
Definition at line 112 of file MR_Reactions.hh.
| ggl::chem::MR_Reactions::MR_Reactions | ( | const Smiles2GraphMap & | smiles2graph, |
| Smiles2GraphMap & | newSmiles2graph, | ||
| Reaction_Container & | reactions, | ||
| const bool | addEachComponent = false, |
||
| const ReactionRateCalculation * | rateCalculation = NULL, |
||
| const AromaticityPerception & | aromaticity = AP_NSPDK("Marvin:general:2013") |
||
| ) |
Construction
| smiles2graph | the initial container that contains all molecules known so far and is fixed |
| newSmiles2graph | the container where all new molecules will be stored in |
| reactions | the Reaction container to fill |
| addEachComponent | if set to true, than all components of the Rules LeftSidePattern are matched to an own target graph copy if two components map to the same target graph |
| rateCalculation | The object to use to calculate a reaction rate for each new reaction created. If not present (==NULL) no reaction rate will be calculated. |
| aromaticity | the aromaticity perception class to be used to correct rings in the product molecules after the rule application was done. |
| ggl::chem::MR_Reactions::MR_Reactions | ( | const Smiles2GraphMap & | smiles2graph, |
| const Smiles2GraphMap & | smiles2graph2, | ||
| Smiles2GraphMap & | newSmiles2graph, | ||
| Reaction_Container & | reactions, | ||
| const bool | addEachComponent = false, |
||
| const ReactionRateCalculation * | rateCalculation = NULL, |
||
| const AromaticityPerception & | aromaticity = AP_NSPDK("Marvin:general:2013") |
||
| ) |
Construction
| smiles2graph | the initial container that contains all molecules known so far and which is not changed |
| smiles2graph2 | second container of the initial molecules known so far |
| newSmiles2graph | the container where all new molecules will be stored in |
| reactions | the Reaction container to fill |
| addEachComponent | if set to true, than all components of the Rules LeftSidePattern are matched to an own target graph copy if two components map to the same target graph, |
| rateCalculation | The object to use to calculate a reaction rate for each new reaction created. If not present (==NULL) no reaction rate will be calculated. |
| aromaticity | the aromaticity perception class to be used to correct rings in the product molecules after the rule application was done. |
|
virtual |
Destruction
|
virtualinherited |
The reported graphs is split into its individual independent components. Each component is forwarded to addMolecule to be implemented by any sub class.
| graph | the graph object to add that encodes one or more molecules |
Implements ggl::Graph_Storage.
|
virtualinherited |
Converts a given molecule graph to SMILES and adds it to the storage container.
| graph | the Graph object to add. |
Implements ggl::chem::GS_chem.
|
protectedvirtual |
Writes a given SMILES and the corresponding molecule graph to the global storage. Furthermore, the given SMILES is added to the products of the currently processed ggl::chem::Reaction.
| SMILES | the SMILES string produced |
| molGraph | the molecule graph that corresponds to the given SMILES |
Reimplemented from ggl::chem::GS_SMILES_MOLp< Smiles2GraphMap >.
|
virtual |
Reports a match. The match is encoded using a vector. The length of the vector corresponds to the number of vertices in the pattern and position i encodes the matched position of pattern node i in the target graph (an instance of sgm::Graph_boostV_p).
| pattern | the pattern graph that was searched for. NOTE: HAS TO BE AN INSTANCE OF ggl::chem::LeftSidePattern !!! |
| target | the graph the pattern was found within. NOTE: HAS TO BE AN INSTANCE OF sgm::Graph_boostV_p !!! |
| match | contains the indices of the matched pattern nodes in the target graph. match[i] corresponds to the mapping of the ith vertex in the pattern graph. |
Implements sgm::Match_Reporter.
|
protected |
if set to true, than all components of the Rules LeftSidePattern are matched to an own target graph copy if two components map to the same target graph
Definition at line 136 of file MR_Reactions.hh.
|
protected |
Definition at line 131 of file MR_Reactions.hh.
|
protected |
Definition at line 125 of file MR_Reactions.hh.
|
protected |
the checker used to correct all molecules produced by the Ruler; it will apply an AP_NSPDK aromaticity perception instance.
Definition at line 140 of file MR_Reactions.hh.
|
protected |
Definition at line 147 of file MR_Reactions.hh.
|
protected |
Definition at line 128 of file MR_Reactions.hh.
|
protected |
the Match_Reporter utilized to apply the rule and to produce the resulting molecules that will be stored in the reaction
Definition at line 144 of file MR_Reactions.hh.
|
protectedinherited |
the map where each SMILES string is mapped to the represented molecule
Definition at line 186 of file GS_SMILES.hh.
|
protectedinherited |
Definition at line 187 of file GS_SMILES.hh.
|
protectedinherited |
Definition at line 188 of file GS_SMILES.hh.