Info
Welcome to the 35th TBI Winterseminar in Bled, exploring Computational Biology, Computational Mathematics, Theoretical Biology, Bioinformatics, Biological Networks.
Time
Sunday, 09 Feb 2020 - Saturday, 15 Feb 2020
Registration Fee
300,- € (double room) / 400,- € (single room)
Participants
- Institute of Theoretical Chemistry (University of Vienna)
- Department of Bioinformatics (University of Leipzig)
- Chair of Bioinformatics (University of Freiburg)
- Center for non-coding RNA in Technology and Health (RTH) (University of Copenhagen)
- RNA Bioinformatics and High-Throughput Analysis Group (Friedrich Schiller University Jena)
- and guests (by invitation only).
Location(s)
Villa Plemelj
Prešernova 39
4260 Bled, Slovenia
Phone: +386 4 5743 023
GPS Coordinates: 46.371121,14.108504
Prešernova 39
4260 Bled, Slovenia
Phone: +386 4 5743 023
GPS Coordinates: 46.371121,14.108504
How to get there
From Ljubljana airport you have the following options to get to the Winterseminar:
From Ljubljana airport you have the following options to get to the Winterseminar:
- take a regular taxi (approx. 50 €)
- public bus transfer, first to Ljubljana city and from there to Bled
- Airport Shuttle ZUP prevozi (special rates for groups)
- New airport public bus to Bled (recommended, cheap and run every hour)
Program
Arrival and Get Together at the Villa
9 Feb 20205pmVilla Plemlj
Opening Session
10 Feb 20204pmHotel Astoria
Graph Theory
11 Feb 20204pmHotel Astoria
Bioinformatics II
8pmHotel Astoria
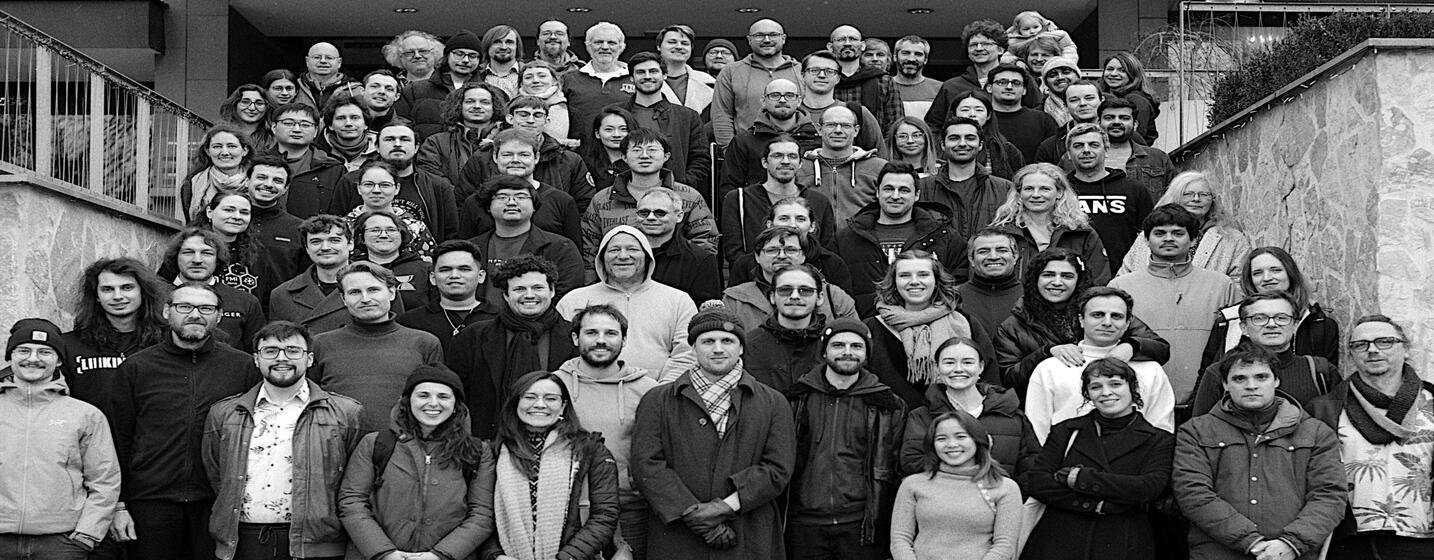
Group Photo
12 Feb 20203:30pmHotel Astoria (Entrance)
RNA I
12 Feb 20204pmHotel Astoria
Transcriptomics
14 Feb 20204pmHotel Astoria
Photos
Further Information
Find detailed information here.
Registered Participants
| Name (headcount: 54) | Affiliation | ||
|---|---|---|---|
| Ronny | Lorenz | ronny (at) tbi.univie.ac.at | TBI Wien |
| Peter F. | Stadler | studla (at) bioinf.uni-leipzIg.de | Bioinformatik Uni Leipzig |
| Yann | Ponty | yann.ponty (at) lix.polytechnique.fr | CNRS/Ecole Polytechnique |
| Emanuel | Barth | emanuel.barth (at) uni-jena.de | FSU Jena |
| Kevin | Lamkiewicz | kevin.lamkiewicz (at) uni-jena.de | FSU Jena |
| Florian | Mock | florian.mock (at) uni-jena.de | FSU Jena |
| Marie | Lataretu | marie.lataretu (at) uni-jena.de | FSU Jena |
| Muriel | Ritsch | anne.muriel.christin.ritsch (at) uni-jena.de | FSU Jena |
| Dominik | Otto | dominik.otto (at) izi.fraunhofer.de | Fraunhofer IZI |
| Julia | Wielach | a01447400 (at) unet.univie.ac.at | TBI Wien |
| Bojana | Ristivojcevic | bojana.risti (at) hotmail.com | TBI Wien |
| Natasha | Jorge | nitandressa (at) gmail.com | Bioinformatik Uni Leipzig |
| Lucas | Vieira | lucas (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| Sebastian | Krautwurst | sebastian.krautwurst (at) uni-jena.de | FSU Jena |
| Sarah | Krautwurst | sarah.krautwurst (at) uni-jena.de | FSU Jena |
| Daniel | Desirò | daniel.desiro (at) uni-jena.de | FSU Jena |
| Christoph | Flamm | xtof (at) tbi.univie.ac.at | TBI Wien |
| Carsten R. | Seemann | carsten (at) bioinf.uni-leipzig.de | MPI MIS Leipzig |
| Ivo | Hofacker | ivo (at) tbi.univie.ac.at | TBI Wien |
| Jakob Lykke | Andersen | jlandersen (at) imada.sdu.dk | SDU Odense |
| Maximilian | Arlt | maximilian.arlt (at) uni-jena.de | FSU Jena |
| Sebastian | Will | will (at) tbi.univie.ac.at | TBI Wien |
| Lisa-Marie | Barf | lisa-marie.barf (at) uni-jena.de | FSU Jena |
| Franziska | Hufsky | franziska.hufsky (at) uni-jena.de | FSU Jena |
| Marcel | Rieger | franziska.hufsky (at) uni-jena.de | family member |
| Marik | Hufsky | franziska.hufsky (at) uni-jena.de | family member |
| Carmen | Bruckmann | carmen.bruckmann (at) ufz.de | UFZ Leipzig |
| Hua-Ting | Yao | hua-ting.yao (at) polytechnique.edu | Ecole Polytechnique/McGill University |
| Christiane | Gärtner | christiane (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| John | Anders | johnanders (at) posteo.de | Bioinformatik Uni Leipzig |
| Mikkel | Pilegaard | mipil15 (at) student.sdu.dk | SDU Odense |
| Tobias | Hagemann | th70tylu (at) studserv.uni-leipzig.de | Bioinformatik Uni Leipzig |
| Stephan | Bernhart | berni (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| Sven | Findeiß | sven (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| Sarah | Berkemer | bsarah (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| Sarah | von Löhneysen | lsarah (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| David | Schaller | sdavid (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| Angel | Rodriguez | aerfangel (at) gmail.com | Bioinformatik Uni Leipzig |
| Maria | Waldl | maria (at) tbi.univie.ac.at | TBI Wien |
| Irene K. | Beckmann | irene (at) tbi.univie.ac.at | TBI Wien |
| Manja | Marz | manja (at) uni-jena.de | FSU Jena |
| Freja Maj | Kelleris | freja (at) rth.dk | RTH Copenhagen |
| Nicolas | Wieseke | wieseke (at) informatik.uni-leipzig.de | SIKS Uni Leipzig |
| Veerendra | Gadekar | veer (at) rth.dk | RTH Copenhagen |
| Rolf | Backofen | backofen (at) informatik.uni-freiburg.de | Uni Freiburg |
| Vivian | Brandenburg | Vivian.Brandenburg (at) rub.de | RUB Bochum |
| Bruno | Sargueil | bruno.sargueil (at) parisdescartes.fr | CNRS |
| Mehmet | Tekman | tekman (at) informatik.uni-freiburg.de | Uni Freiburg |
| Philipp | Honegger | philipp.honegger (at) univie.ac.at | MDY Wien |
| Josef | Leydold | josef.leydold (at) wu.ac.at | WU Wien |
| Stefanie | Kehr | steffi (at) bioinf.uni-leipzig.de | Bioinformatik Uni Leipzig |
| Anahy | Santiago | jpscw (at) hotmail.com | Bioinformatik Uni Leipzig |
| Celia | Diezel | celia.diezel (at) uni-jena.de | FSU Jena |
| Nino | Basic | nino.basic (at) famnit.upr.si | University of Primorska, Koper |